'CAinterprTools': R package for visual aid to Correspondence Analysis interpretation
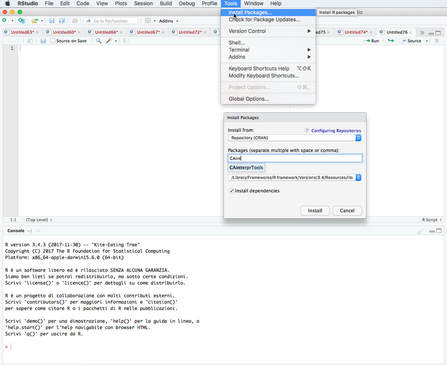
Some of the features of the R script for CA (described in this site) have been turned into an R package. As of July 2018, the package is available from CRAN and can therefore easily installed into R. For other info, and for download statistics, see the RDocumentation website.
Besides implementing some of the features of my CA script, the package dramatically expands the available facilities to get a visual aid to the interpretation of the results of Correspondence Analysis. Among other things, the package allows to calculate the significance of the CA dimensions and of the total inertia by means of a permutation test.
The package is also described in an article of mine published in Elsevier's SoftwareX journal (LINK).
If you want to cite my package, you may use the following format:
Alberti G, CAinterprTools: An R package to help interpreting Correspondence Analysis’ results, SoftwareX, Volumes 1–2, September 2015, Pages 26-31, ISSN 2352-7110, http://dx.doi.org/10.1016/j.softx.2015.07.001.
Please note that the package has been improved several times since 2015. A description of the facilities it provides (as of version 1.0.0) can be found in the PDF below.
Some of the features of the R script for CA (described in this site) have been turned into an R package. As of July 2018, the package is available from CRAN and can therefore easily installed into R. For other info, and for download statistics, see the RDocumentation website.
Besides implementing some of the features of my CA script, the package dramatically expands the available facilities to get a visual aid to the interpretation of the results of Correspondence Analysis. Among other things, the package allows to calculate the significance of the CA dimensions and of the total inertia by means of a permutation test.
The package is also described in an article of mine published in Elsevier's SoftwareX journal (LINK).
If you want to cite my package, you may use the following format:
Alberti G, CAinterprTools: An R package to help interpreting Correspondence Analysis’ results, SoftwareX, Volumes 1–2, September 2015, Pages 26-31, ISSN 2352-7110, http://dx.doi.org/10.1016/j.softx.2015.07.001.
Please note that the package has been improved several times since 2015. A description of the facilities it provides (as of version 1.0.0) can be found in the PDF below.
Have you found this website helpful? Consider to leave a comment in this page.